 A.C. Gerstein, A. Kuzmin & S.P. Otto (2014) Loss of heterozygosity facilitates passage through Haldane's sieve for Saccharomyces cerevisiae undergoing adaptation. Nature Communications. This one has been a long time coming. It's the last chapter of my dissertation and the paper I'm the most proud of so far. This paper documents what I set out to study during my PhD, the dominance of beneficial mutations. It is also a testament to pushing through even when results are confusing and unexpected. I spent about six months doing experiments that I couldn't make sense of. I have previously published on a set of mutations I acquired in haploids that are resistant to nystatin, an antifungal drug. We identified 20 unique mutations among 30 lines that are clustered into four genes in the ergosterol biosynthesis pathway [Gerstein et. al (2012), Genetics]. From the initial haploid lines I created homozygous and heterozygous lines for all 20 unique mutations. The homozygous story is interesting in its own right, I unexpectedly found that the mutations have different effect sizes in haploids and homozygous diploids [Gerstein (2013), Biology Letters]. Truthfully, however, the reason why I acquired the mutations in the first place was to study their fitness effects in a heterozygous background. Logically, the heterozygous mutations should have fitness values the same as wildtype (i.e., recessive mutations), the same as the homozygous mutants (dominant mutations) or somewhere in between (and maybe a small number would have fitness greater than the homozygotes, i.e., overdominance). What I found, instead, was that my fitness assays were all over the place for the heterozygotes, they simply weren't behaving like I expected and I thought that I was doing something wrong (even though these are experiments I had done many many times before). Turns out that biology is craftier than I gave it credit - my heterozygotes individuals simply weren't always staying heterozygous! Instead, the stochasticity in my assays was showing me that loss of heterozygosity (LOH) was frequently happening within the populations, during the short (~72h) timeframe of my experiments. In hindsight, this shouldn't have been surprising. LOH is a fairly common mutation, and it provides a really quick way for a recessive heterozygous mutation to behave like a dominant mutation - just ditch the wildtype allele in favour of a second beneficial mutant allele. What this means, as an evolutionary biologist, is that recessive beneficial mutations, which are predicted to play a minor role in evolution because they should get sieved out of the population (as shown above) can bypassed the sieve (known in the theoretical literature as Haldane's sieve) by LOH. This will be particularly important for asexual diploids, where it is otherwise difficult for recessive heterozygous mutations to have a fitness effect. I'm also proud of this paper because we fought for it. We sent it to six different journals before this one - it was reviewed and rejected by an editor at one journal and had a full review and got rejected at a second. Then, rather than getting frustrated and sending it to a lower tier of journal, with the encouragement of my advisor I stripped it down and majorly streamlined it. We then sent it to Nature Communications, and after some thoughtful reviews, and an additional experiment that I think adds a lot, it should be online there in the upcoming week. 14.04.24. UPDATE: Apparently someone at SGD also thought the biology in this paper was neat - our paper is featured in a blog post! 14.07.11 UPDATE: And so did Stephen Wright at F1000Prime - recommended as an exceptional read. 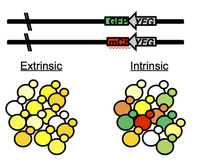 M.Z. Anderson, A.C. Gerstein, L. J. Wigen, J.A. Baller, J. Berman Silencing is noisy: Population and cell level noise in telomere-adjacent genes is dependent on telomere position and Sir2. PLoS Genetics. This paper is something I began working on at the start of my postdoc, with Matt Anderson, another postdoc in the Berman Minnesota lab who has now moved on to Richard Bennett's lab at Brown University. I came on board as the experimental design consultant/ statistician/graph-maker. Matt had collected an impressive amount of data and I came in to statistically analyze it (/ recollect it with proper controls done on the same day :-) and make piles of R graphs. I got to learn about chromatin and Sir2 and telomeres, and I got to teach Matt about bootstrapping, experimental design and the virtues of plotting in R.
14 Comments
|
Archives
September 2018
Categories |
 RSS Feed
RSS Feed