|
I was thrilled to get to talk at the American Naturalist stand alone meeting in a symposium to highlight influential papers that have been published in the journal. When I saw the call for talks last year I immediately knew that someone had to talk about the Lenski et al. 1991 paper that is typically credited as introducing experimental evolution to the larger eco-evo community (note that microbiologists had already been doing these types of experiments since at the least the 1950s, but somewhere along the line the two groups largely stopped talking to each other). It was a fun talk to put together, since the goal was to highlight the influence of the paper on the field and on my own work. It turns out that wanting to make sure you accurately represent the impact of the work of one of your scientific heros is kind of stressful! I think it worked out though ... The meeting itself was lovely, I had embargoed myself from going to ecology and evolution meetings as a postdoc until this past year and although I learned a LOT going to fungal genetics, cell biology and molecular biology meetings, eco-evo will always be home. Sidenote: I think there's still a lack of both microbiology and mycology in the field, and although I am in the strangely (but perhaps not so unique) precarious position of having no idea where I'm going to end up in the next months and years, I do think work on fungal microbes has important things to tell us about both ecology and evolution.
So clearly this blog thing didn't really take. But no time like the present to try again? I'm off tomorrow for Portland for the Evolution Meeting. The last time I was at the Evolution meeting was when it was the super meeting in Ottawa in 2012. That was a crazy time for me, I defended my PhD thesis about 10 days afterwards! It was also the meeting where I presented the culmination of five years of really hard PhD work and was (co-)awarded the Hamilton award for best student talk.
As a postdoc I have been focused really heavily on learning as much cellular, molecular and fungal biology as possible, and so have purposefully attended meetings where I could advance that knowledge. Last fall I was feeling pretty down and isolated and decided that I wanted to go back to Evolution to 'come home'. The Evolution Meetings played a huge role in my graduate experience. The Evolution meeting in Fort Collins Colorado (in 2004!) was my first real science meeting. I completely remember how awed I was that so many people studied evolution. I've volunteered to serve as a peer mentor to two undergraduate students and I'm really looking forward to reliving that experience through them. I'm also really excited about my talk (Sunday morning, 9am in the Convergent/Parallel Evolution session in the Oregon Ballroom). I'm going to be presenting on work that I started in my first year of graduate school and have been slowly working on ever since. It's also what I presented at the Evolution Meeting in Alaska in 2005, my first meeting talk! And for the first time in awhile, my talk is finished before I get on the airplane. Win!  A.C. Gerstein, A. Kuzmin & S.P. Otto (2014) Loss of heterozygosity facilitates passage through Haldane's sieve for Saccharomyces cerevisiae undergoing adaptation. Nature Communications. This one has been a long time coming. It's the last chapter of my dissertation and the paper I'm the most proud of so far. This paper documents what I set out to study during my PhD, the dominance of beneficial mutations. It is also a testament to pushing through even when results are confusing and unexpected. I spent about six months doing experiments that I couldn't make sense of. I have previously published on a set of mutations I acquired in haploids that are resistant to nystatin, an antifungal drug. We identified 20 unique mutations among 30 lines that are clustered into four genes in the ergosterol biosynthesis pathway [Gerstein et. al (2012), Genetics]. From the initial haploid lines I created homozygous and heterozygous lines for all 20 unique mutations. The homozygous story is interesting in its own right, I unexpectedly found that the mutations have different effect sizes in haploids and homozygous diploids [Gerstein (2013), Biology Letters]. Truthfully, however, the reason why I acquired the mutations in the first place was to study their fitness effects in a heterozygous background. Logically, the heterozygous mutations should have fitness values the same as wildtype (i.e., recessive mutations), the same as the homozygous mutants (dominant mutations) or somewhere in between (and maybe a small number would have fitness greater than the homozygotes, i.e., overdominance). What I found, instead, was that my fitness assays were all over the place for the heterozygotes, they simply weren't behaving like I expected and I thought that I was doing something wrong (even though these are experiments I had done many many times before). Turns out that biology is craftier than I gave it credit - my heterozygotes individuals simply weren't always staying heterozygous! Instead, the stochasticity in my assays was showing me that loss of heterozygosity (LOH) was frequently happening within the populations, during the short (~72h) timeframe of my experiments. In hindsight, this shouldn't have been surprising. LOH is a fairly common mutation, and it provides a really quick way for a recessive heterozygous mutation to behave like a dominant mutation - just ditch the wildtype allele in favour of a second beneficial mutant allele. What this means, as an evolutionary biologist, is that recessive beneficial mutations, which are predicted to play a minor role in evolution because they should get sieved out of the population (as shown above) can bypassed the sieve (known in the theoretical literature as Haldane's sieve) by LOH. This will be particularly important for asexual diploids, where it is otherwise difficult for recessive heterozygous mutations to have a fitness effect. I'm also proud of this paper because we fought for it. We sent it to six different journals before this one - it was reviewed and rejected by an editor at one journal and had a full review and got rejected at a second. Then, rather than getting frustrated and sending it to a lower tier of journal, with the encouragement of my advisor I stripped it down and majorly streamlined it. We then sent it to Nature Communications, and after some thoughtful reviews, and an additional experiment that I think adds a lot, it should be online there in the upcoming week. 14.04.24. UPDATE: Apparently someone at SGD also thought the biology in this paper was neat - our paper is featured in a blog post! 14.07.11 UPDATE: And so did Stephen Wright at F1000Prime - recommended as an exceptional read. 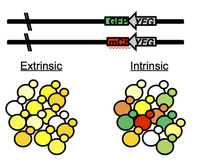 M.Z. Anderson, A.C. Gerstein, L. J. Wigen, J.A. Baller, J. Berman Silencing is noisy: Population and cell level noise in telomere-adjacent genes is dependent on telomere position and Sir2. PLoS Genetics. This paper is something I began working on at the start of my postdoc, with Matt Anderson, another postdoc in the Berman Minnesota lab who has now moved on to Richard Bennett's lab at Brown University. I came on board as the experimental design consultant/ statistician/graph-maker. Matt had collected an impressive amount of data and I came in to statistically analyze it (/ recollect it with proper controls done on the same day :-) and make piles of R graphs. I got to learn about chromatin and Sir2 and telomeres, and I got to teach Matt about bootstrapping, experimental design and the virtues of plotting in R.  Hello world. That's what you're suppose to write the first time you start one of these things, right? I'm sitting in my (two bedroom + dining room + kitchen + backyard) Minneapolis apartment, drinking coffee, listening to Asaf Avidan. I've decided that I'm done with iWeb, so here I am at Weebly after some inspiration from the O'Conner lab at UBC. And as these things go I somehow ended up with a blog page. I'm almost nine month into my bi-continental postdoc. I've accumulated enough air miles in the year since finishing my PhD that I'm going to be a Sky Alliance Platinum member next year. I've got one more week in Minneapolis before another quick week in Winnipeg until I head back to Tel Aviv until Christmas. Hopefully next year will be more stable (primarily in Minneapolis, a totally great city that I haven't really been able to spend enough time in to settle into yet). I did manage to acquire a new mode of commuting (on the left) to fully enjoy living here though. |
Archives
September 2018
Categories |
 RSS Feed
RSS Feed